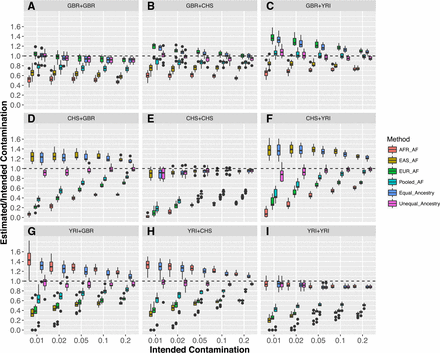
Comparison of different models to estimate contamination rates. Horizontal (x) axis shows intended contamination rate, vertical (y) axis shows the ratio of estimated to intended contamination rates. Each color represents different models to estimate contamination rates. EUR_AF, EAS_AF, and AFR_AF represent original verifyBamID using European, East Asian, and African allele frequencies across the continental population using the 1000 Genomes data. Pooled_AF represents the original verifyBamID using aggregated allele frequencies across all 2504 individuals in the 1000 Genomes Project. Equal_Ancestry represents the verifyBamID2 assuming that intended and contaminating samples belong to the same population. Unequal_Ancestry represents verifyBamID2 allowing different genetic ancestry between intended and contaminating sample (recommended setting). Each panel (A–I) represents different combinations of intended (row) and contaminating (column) populations, in the order of GBR, CHS, and YRI.











