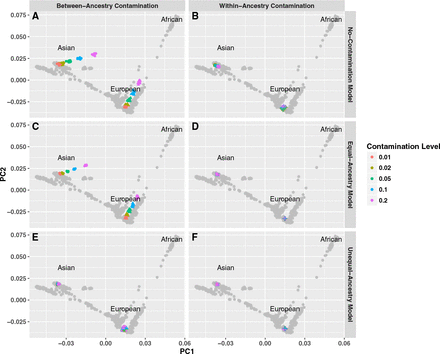
Impact of DNA sample contamination on the estimation of genetic ancestry. Each point represents a sample. Each gray point represents reference (HGDP) sample and its PCA coordinates, similar to Figure 2. Each colored point represents in silico–contaminated samples across various contamination rates and populations. In panels A, C, and E, European (GBR) and East Asian (CHS) samples are contaminated with African (YRI) samples at different contamination rates (i.e., between-ancestry contamination). In panels B, D, and F, European (GBR) and East Asian (CHS) samples are contaminated with another sample in the same population (i.e., within-ancestry contamination). Different colors represent different contamination rates ranging from 1% to 20%. Upper panels (A,B) show verifyBamID2 estimates without modeling contamination; middle panels (C,D), verifyBamID2 estimates under the assumption that intended and contaminating populations are identical (i.e., equal-ancestry model); lower panels (E,F), verifyBamID2 estimates under the assumption that intended and contaminating populations can be different (i.e., unequal-ancestry model).











