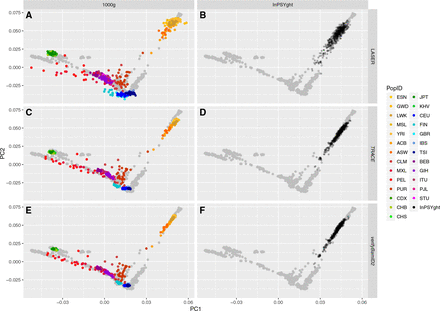
Evaluation of estimated genetic ancestry coordinates, in the absence of contamination, between TRACE, LASER, and verifyBamID2 on samples from the 1000 Genomes low-coverage genome (n = 500, diverse ancestry) sequence data (A,C,E) and from the InPSYght deep genome (n = 500, African-Americans) sequence data (B,D,F). Panels A and B show results from TRACE, C and D from LASER, and E and F from verifyBamID2 (assuming no contamination). Each point represents a sample and each color represents a population ancestry, with the exception that gray points represent PCA coordinates of reference (HGDP) samples.











