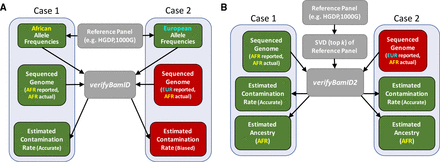
Overview of verifyBamID and verifyBamID2 software tools. (A) verifyBamID takes aligned sequence reads (in BAM format) and known variant sites annotated with population allele frequencies (in VCF format) to estimate DNA contamination rates. When allele frequencies are correctly specified, the estimated DNA contamination rates are expected to be accurate (green boxes). However, when the allele frequencies are misspecified (e.g., due to incorrect self-reported ancestry), the estimates of DNA contamination rates may be biased (red boxes). (B) verifyBamID2 takes aligned sequence reads (in BAM/CRAM format) and top k singular value decomposition (i.e., PCs and SNP loadings) to estimate the genetic ancestries and contamination rates together. Because verifyBamID2 does not rely on self-reported ancestry, even if ancestry of sample is misspecified or unknown (red box), the estimated contamination rates will be unbiased (green box). In addition, genetic ancestries are also estimated in PC coordinates, adjusting for potential contamination.











