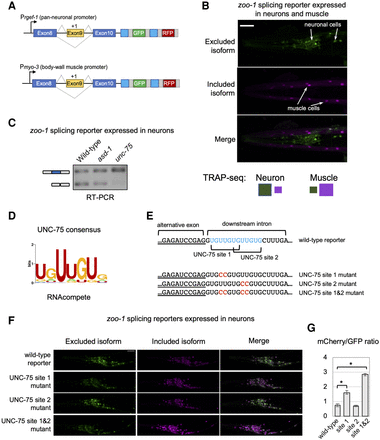
UNC-75/CELF represses inclusion of an exon by binding to cis elements overlapping a 5′ splice site. (A) Schematic of two-color reporters used for experiments. (Prgef-1) Promoter driving pan-neuronal expression; (Pmyo-3) promoter driving body-wall muscle expression. (B) Fluorescence microscopy images of zoo-1 two-color reporter monitoring splicing of exon 9 in both neurons and muscle cells. Neurons display increased exon skipping (green), whereas muscle cells primarily include the alternative exon (purple). Scale bar, 20 μm. (C) RT-PCR assay monitoring zoo-1 two-color splicing reporter expressed only in neurons in wild-type animals (lane 1), asd-1 mutants (lane 2), and unc-75 mutants (lane 3). Primers are designed to amplify both exon 9 included (top band) and excluded (bottom band) isoforms. (D, left) UNC-75 consensus motif derived from RNAcompete data (Ray et al. 2013). (E, top) Schematic of 3′ end of zoo-1 exon 9 (underlined) and flanking downstream intron sequence. Two UNC-75 consensus sequences overlapping the 5′ splice site are highlighted in blue. (Bottom) Reporters with mutated UNC-75 consensus sequences (substitutions in red). (F) Fluorescence microscopy images of wild-type and mutant two-color zoo-1 reporters as described in E. Scale bar, 20 μm. (G) Bar graph displaying mCherry/GFP fluorescence intensity ratios for each zoo-1 splicing reporter. Each bar represents mean ratio from five individual animals. Error bars, ± 1 SD from the mean value. (*) P-value < 0.01, Kruskal–Wallis rank sum test.











