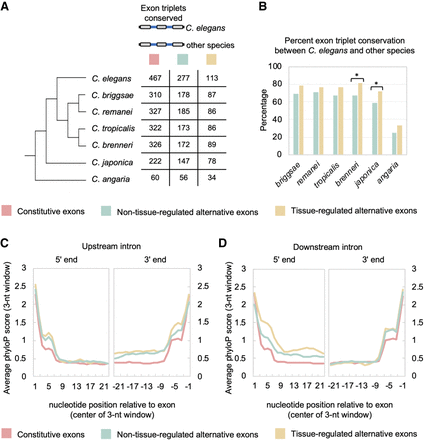
Alternative and tissue-regulated splicing are associated with increased conservation patterns. (A, left) Phylogenetic tree of Caenorhabditis species used in alignments. (Right) Splicing events were separated into three classes of exon triplets: constitutive exons, non-tissue-regulated alternative exons, and tissue-regulated alternative exons. Table shows number of exon triplets in each group and their conservation across the phylogeny. (B) Graph of percentage of conserved exon triplets for non-tissue-differential alternative exons (light blue bars) and tissue-differential alternative exons (gold bars) between C. elegans and other Caenorhabditis species of varying divergence. (*) P-value < 0.02, Fisher's exact test. (C) Plot of average phyloP score (rolling 3-nt average) over first 23 nt (left) and last 23 nt (right) of the upstream intron flanking the internal exon of the exon triplets described in panel A. (D) Same as in C, but plots of the average phyloP score for the downstream intronic regions.











