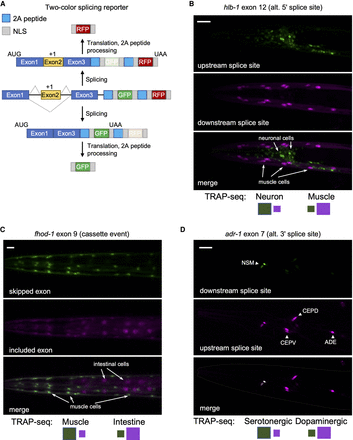
Validation of TRAP-seq measurements with two-color splicing reporters. (A) Schematic of the two-color splicing reporter architecture. A minigene, comprising an alternatively spliced cassette exon (yellow) with flanking introns and flanking constitutive exons (blue), is cloned upstream of GFP and mCherry (RFP) ORFs. The alternative exon is engineered to include an extra nucleotide (+1) to shift reading frame. The ORFs differ in reading frame resulting in the expression of RFP upon inclusion of the cassette exon, whereas skipping of the exon leads to the expression of GFP. Cleavage at the 2A peptides and nuclear localization sequences (NLSs) ultimately leads to free fluorescent reporter proteins to accumulate in the nucleus. (B) Fluorescence microscopy of splicing reporter expressed in neurons and muscle cells monitoring a splicing event in the hlb-1 gene. Top labels indicate gene, alternatively spliced exon, and splicing class; bottom labels indicate predominant color expected in reporter for a particular tissue (larger-size green or purple square) from measured PSI values from TRAP-seq data. Scale bar, 20 μm. (C) Same as in B, with splicing reporter expressed in intestine and muscle cells monitoring splicing event in fhod-1 gene. Scale bar, 20 μm. (D) Same as in B and C, with splicing reporter expressed in dopaminergic (CEP and ADE) and serotonergic (NSM) neurons monitoring splicing event in adr-1 gene. Scale bar, 20 μm.











