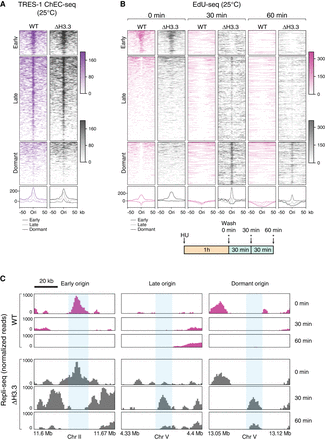
Replication origin dynamics are altered in H3.3-null mutants. (A) Heatmaps (top) and average plots (bottom) of TRES-1 ChEC-seq signal at replication origins, for wild-type (WT, purple) and H3.3-null mutant (Δ H3.3, gray) worms grown at 25°C. Signal is shown for a 50-kb window around each origin. (B,C) EdU-seq time course, for WT (pink) and Δ H3.3 (gray) worms grown at 25°C. (B) Heatmaps (top) and average plots (bottom) of EdU-seq signal at 0, 30, and 60 min at all origins. Signal (normalized reads) is shown for a 50-kb window around each origin. The cartoon shows the experimental protocol, in which cells were incubated 1 h with HU, released from the HU block, and treated with EdU and HU at 0, 30, or 60 min (indicated by the asterisks). (C) Genome browser views of EdU-seq signal at 0, 30, and 60 min, showing fork progression at representative examples of early, late, and dormant origins.











