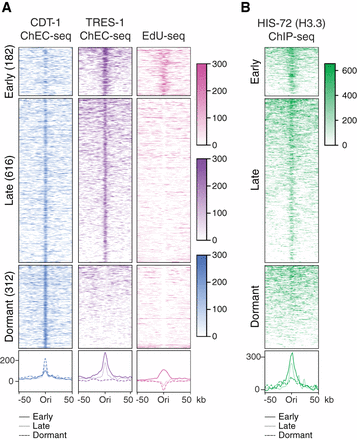
Classification of replication origins in C. elegans embryos. (A) Heatmaps (top) and average plots (bottom) of CDT-1 ChEC-seq (blue), TRES-1 ChEC-seq (purple), and EdU-seq (pink) signal at replication origins. Origins were identified by peak calling using the CDT-1 ChEC-seq, TRES-1 ChEC-seq, and EdU-seq data sets obtained from worms grown at 20°C and classified into early, late, and dormant origins through unsupervised clustering. Signal (normalized reads) is shown for a 50-kb window around each origin. (B) Heatmaps (top) and average plots (bottom) of HIS-72 (H3.3) ChIP-seq signal (Delaney et al. 2019) at replication origins, as in A.











