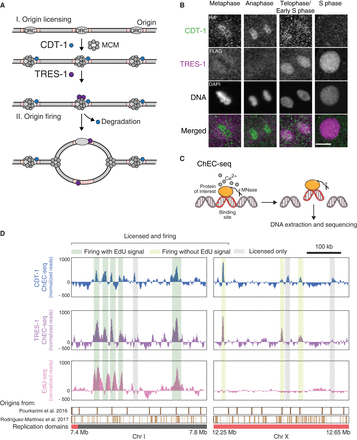
Identification of replication origins in C. elegans embryos. (A) Schematic description of the roles of CDT-1 and TRES-1 in replication origin firing. CDT-1 is required for the licensing of all origins. TRES-1 is recruited only to origins that fire. (B) Localization of HA::CDT-1 (green) and FLAG::TRES-1 (magenta) during the cell cycle. Immunofluorescence images using anti-HA and anti-FLAG antibodies and DAPI are shown. Scale bar represents 5 µm. (C) Schematic description of ChEC-seq. The protein of interest is fused with MNase. Upon activation with calcium in purified nuclei, MNase cleaves and releases DNA fragments at the binding sites of the fusion proteins. These small fragments are isolated and sequenced. (D) Representative genome browser views of CDT-1 ChEC-seq (blue), TRES-1 ChEC-seq (purple), and EdU-seq (pink) signal. Identified origins are highlighted in gray (licensed only) or dark and light green (licensed and firing with and without EdU signal, respectively). Domains of late (red) and early (gray) replication and positions of replication origins identified in previous studies are shown below the genome browser (Pourkarimi et al. 2016; Rodríguez-Martínez et al. 2017).











