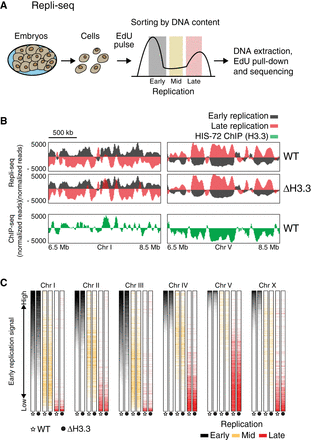
Replication timing is unchanged in the absence of H3.3. (A) Schematic description of Repli-seq. Embryonic cells were dissociated, exposed to an EdU pulse for 5 min, and sorted according to their DNA content. EdU-labeled DNA was sequenced and mapped to the genome. (B) Representative genome browser views of Repli-seq and H3.3 ChIP-seq. Repli-seq signal is shown for wild-type (WT) and H3.3-null mutant (Δ H3.3) worms on regions of Chromosomes I and V. Early S phase is shown in black and late S phase in red. HIS-72 (H3.3) ChIP-seq signal is shown in green for the same regions. ChIP-seq data from Delaney et al. (2019). (C) Color-coded replication timing for each chromosome. Repli-seq signal from early (black), mid (orange), and late (red) S phase for WT and Δ H3.3. Data for each chromosome were sorted according to the signal of early S phase in WT.











