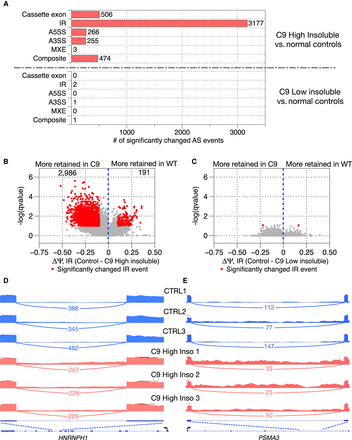
Widespread, elevated intron retention is observed in C9high patient brain samples. (A) Number of significantly differentially spliced AS events in six AS categories comparing C9high patient samples with normal controls and C9low patient samples with normal controls, respectively. (B) Volcano plot showing the magnitude and direction of changes in intron retention for a total of 3177 changed intron retention events between C9high patient samples and normal controls. X-axis shows the difference of intron-retained isoform levels between normal controls and C9high patient samples. (C) Volcano plot showing the magnitude and direction of changes in intron retention between C9low patient samples and normal controls. (D,E) Sashimi plot and genome browser shots for two significantly elevated intron retention events in two gene transcripts, HNRNPH1 and PSMA3. Exon coverage from RNA-seq data is shown in three normal control samples (blue) and three C9high samples (red); arcs represent splice junctions identified from the RNA-seq data and the number of uniquely mapped RNA-seq reads mapped to the junctions are shown across the arc; human annotation (hg38) of the transcripts is shown at the bottom. The black arrow indicates the direction of the promoter. The dotted lines indicate the region of the transcript that is enlarged to highlight the retained intron region.











