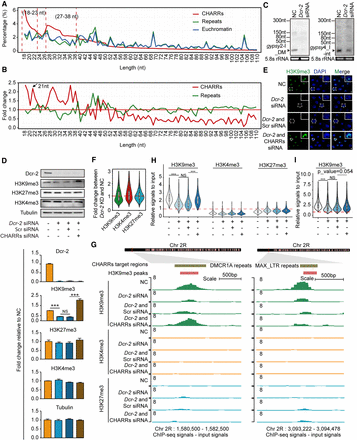
Trans-acting repeat-derived RNAs for heterochromatin maintenance. The length distribution in 1-nt bins based on the combined ChAR-seq data set (A), and at each bin, reads from CHARRs (red) or other repeats (green) were normalized against those from euchromatin RNAs (B). (C) Confirmation of Dcr-2-dependent expression of gypsy2-I_DM (left) and gypsy4_I-int (right) expression by northern blotting analysis. (D) Western blotting analysis of Dcr-2, H3K9me3, H3K27me3, H3K4me3, and Tubulin in S2 cells in response to siRNA-mediated knockdown of Dcr-2 and rescue with a transfected pool of CHARR-derived synthetic siRNAs. Quantified data are shown below as fold-change (FC) relative to mock-treated cells (NC) in lane 1. Data are presented as mean ± SEM (n = 3 biological replicates). (***) P < 0.001, (NS) not significant (unpaired Student's t-test). (E) H3K9me3 detected by immunocytochemistry in S2 cells treated with different combinations of siRNAs, as indicated. (Green) H3K9me3 signals, (blue) DAPI. Scale bar, 2 μm. (F) Violin plot for fold-change of H3K9me3, H3K4me3, and H3K27me3 signals on targeted peaks in response to Dcr-2 knockdown. (KD) Knockdown. (G) H3K9me3 ChIP-seq signals on representative genomic loci, one corresponding to a CHARR-target locus (left) and a non-CHARR-target locus (right) in response to Dcr-2 knockdown, complemented with either scrambled or derived siRNAs. ChIP-seq signals for euchromatin (H3K4me3) or facultative heterochromatin (H3K27me3) were shown for comparison. (H,I) H3K9me3 signals on targeted (H) or nontargeted (I, with <50% sequence homology to any of derived siRNAs) regions in response to Dcr-2 knockdown, complemented with either scrambled or derived siRNAs. (***) P < 0.001 (Student's t-test). H3K27me3 and H3K4me3 binding signals were displayed for comparison.











