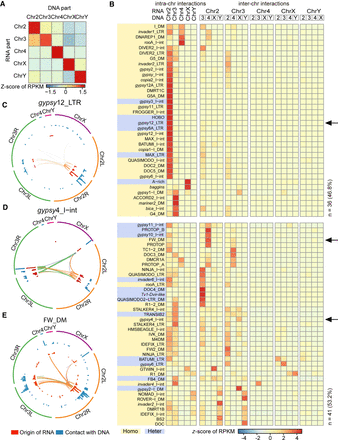
Cis- and trans-acting repeat-derived RNAs on chromatin. (A) Heat map of RNA–DNA interaction scores for 77 CHARRs on the same or different chromosomes. Trans-chromosome interactions are referred to as RNA interacting with DNA in chromosomes other than those from which the RNA is transcribed. (B) Assignment of each of the 77 CHARRs to either intra- or inter-chromosomal interactions in S2 cells and sorting of the data by unsupervised clustering. Top: RNA derived from different chromosomes (first row) and its DNA contacts in the same or different chromosomes (second row). The RNA–DNA interaction intensities were indicated by the z-scores according to the color key at the bottom. Arrows point to the three specific CHARRs gypsy12_LTR, gypsy4_I-int, and FW_DM, as individually illustrated in panels C–E. All CHARRs are either highlighted with yellow or blue to indicate their preferences for homotropic or heterotropic interactions, as described later. (C–E) The origins of three representative CHARRs and their DNA contacts, as shown by the Circos plots for gypsy12_LTR (C), gypsy4_I-int (D), and FW_DM (E). In each of these plots, the inner (red) track indicates the origins of individual RNAs and the outer (blue) track shows where the RNAs interact with DNA in 100-kb DNA windows. The heights of the signals correspond to reads per 100-kb window per millions. Lines specify intra- or inter-chromosomal interactions, with width suggesting the relative interaction levels in each case. The line color indicates the origin of chromatin-interacting RNAs.











