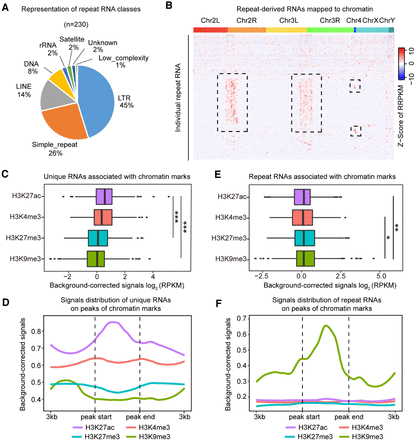
Interaction of distinct RNA classes with eu- versus heterochromatin. (A) Different repeat RNA classes represented by 230 DNA-bound repeat-derived RNAs. (B) Heat map showing the distribution of individual 230 repeat-derived RNAs across the Drosophila genome in S2 cells. (Row) Individual chromatin-associated repeat-derived RNAs, (column) AluI DNA bins, (boxed regions) repeat-derived RNAs that showed preferential binding to constitutive heterochromatin in pericentromeric regions. (C) Association of chromatin marks with background-corrected signals of unique RNAs. (***) P < 0.001 (unpaired Student's t-test). (D) Distribution of unique RNA interaction signals around chromatin mark peaks. (E) Association of chromatin marks with background-corrected signals of repeat-derived RNAs. (*) P < 0.05, (**) P < 0.01 (unpaired Student's t-test). (F) Distribution of repeat RNA interaction signals around chromatin mark peaks.











