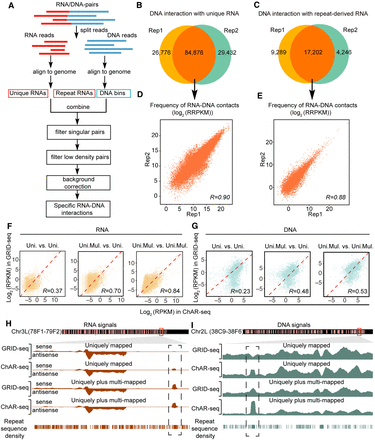
Strategy for assigning multimapped RNA and DNA reads. (A) Schematic presentation of the GRID-seq data processing pipeline. (B,C) Overlapped RNA–DNA mates containing annotated unique (B) or repeat-derived (C) RNA between the two independent GRID-seq libraries. (D,E) Quantitative analysis of commonly identified RNA–DNA mates containing annotated unique RNA (D) or repeat-derived RNA (E) between the two independent GRID-seq libraries. (RRPKM) Read counts per kilobase of RNA per kilobase of DNA per million. (F,G) Comparison between GRID-seq and ChAR-seq with uniquely or uniquely plus multimapped RNA (F) or DNA (G) in repeat-enriched AluI-generated DNA bins. (Uni.) Uniquely mapped reads, (Uni.Mul.) uniquely plus multimapped reads. (H,I) Transcribed RNA signals (H) or RNA-contacted DNA signals (I) obtained with different mapping strategies on representative genomic regions.











