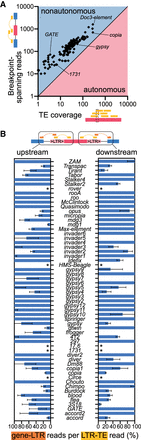
LTR retrotransposon expression is predominantly nonautonomous. (A) Plot showing the average number of reads per nucleotide (x-axis) and total number of transposon–gene spanning reads (y-axis) for every tested transposon. Number of spanning reads is higher for every transposon. (B) List of all LTR retrotransposons analyzed in mRNA data. LTR-gene spanning reads were identified for every LTR transposon expressed in the midbrain. Numbers represent percentage of reads spanning LTR-gene versus LTR-TE breakpoints. Values are capped at 100%, but some transposons produced more LTR-gene than LTR-TE reads (Supplemental Table S10). Error bars represent standard deviation. (*) Transposon reference sequences did not contain LTR sections for the five transposons.











