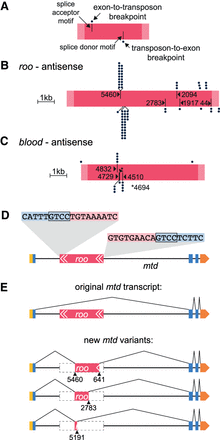
Transposons introduce splice sites at conserved locations. (A) Key for labeling scheme in B and C. Pink bar represents the transposon; light pink ends indicate LTRs, and dark pink indicates the core sequence. Positions of dots above the bar represent sites on the transposon where an upstream exon splice donor (SD) has merged. Every dot represents a different gene. Black lines in the top half of pink bar represent splice-acceptor (SA) motifs in the transposon. Dots below the pink bar indicate location of breakpoints on the transposon that splice to upstream exonic SA sites of different genes. Bars in the lower half indicate SD motifs. (B,C) Representations of antisense roo and blood (to scale), with all breakpoints to SA and SD sites of neighboring genes. The frequently used site on antisense roo at position 5191 is a nonconsensus SD site, lacking the expected GT motif at the immediate breakpoint. The sequence around 5191 resembles a consensus SD motif, although the GT is a GC. Compare TTTGGCAAGTT to motif in Supplemental Figure S5A. (D) Illustration of antisense roo insertion in the mustard (mtd) gene. Only one isoform of mtd is shown. Yellow box represents 5′ UTR, blue boxes are exons, orange box 3′ UTR, pink represents roo transposon with white arrows indicating LTRs. Breakpoint-spanning gDNA reads reveal target site duplication (TSD; inset). (E) Schematic of original mtd transcript and of three new splice isoforms.











