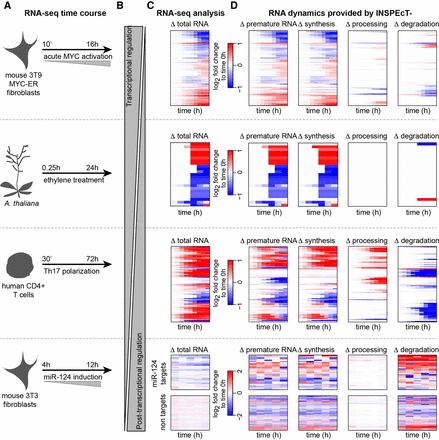
Characterization of time course RNA dynamics: reanalysis of four published data sets with INSPEcT−. (A) Experimental design of the considered RNA-seq time courses. (B) Expected balance between transcriptional and post-transcriptional responses in the different experiments. (C) The typical output of differential RNA-seq analyses: heatmap of differentially expressed genes. (D) The additional information gained by reanalyzing those data with INSPEcT−, which includes the gene-level modulation of premature RNAs, as well as the temporal changes of the kinetic rates of synthesis, processing, and degradation. For the miR-124 data set, reported rates are first-guess estimates, owing to lack of replicates in the time course.











