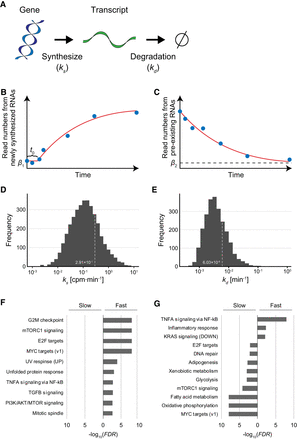
Estimation of RNA synthesis and degradation rates of individual genes. (A) Kinetics model of gene expression. In this model, a transcript was synthesized according to the RNA synthesis rate, ks (CPM/min) and degraded according to the degradation rate, kd (min−1). (B) Synthesis rates were estimated by fitting of the time series of the amount of newly synthesized RNAs from individual genes to a logarithmically increasing curve (see Methods). β1 and t0 indicate the basal value of newly synthesized RNAs and the time delay, respectively. (C) Degradation rates were estimated by fitting of the time series of the amount of pre-existing RNAs from individual genes to an exponentially decreasing curve (see Methods). β2 indicates the basal value of pre-existing RNAs. (D,E) Distribution of estimated RNA synthesis rate (D) and degradation rate (E). (F,G) Gene Set Enrichment Analysis (GSEA) for genes with ks values (F) or with kd values (G).











