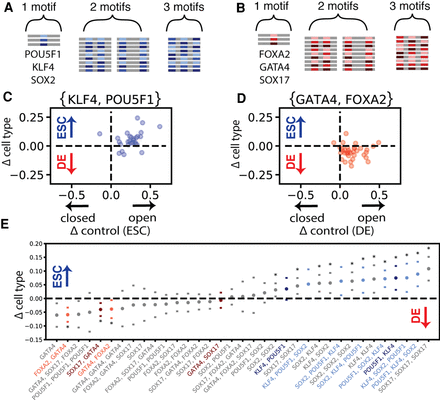
Lineage transcription factor motifs impact chromatin accessibility with preferential spatial ordering. (A) DNA sequence construction from the ESC key transcription factors POU5F1, SOX2, and KLF4. (B) DNA sequence construction from the DE key transcription factors GATA4, SOX17, and FOXA2. (C) Each dot represents a single neutral DNA background sequence that contains one instance of a POU5F1 motif and one instance of a KLF4 motif (two total motif instances per DNA sequence). On the y-axis is the difference between endoderm and ESC accessibility, and on the x-axis is the difference between each DNA sequence and its shuffled control in ESCs. (D) Each dot represents a single neutral DNA background sequence that contains one instance of a GATA4 motif and one instance of a FOXA2 motif (two total motif instances per DNA sequence). On the y-axis is the difference between endoderm and ESC accessibility, and on the x-axis is the difference between each DNA sequence and its shuffled control in DE cells. (E) All motif orderings that had significant accessibility relative to random shuffled DNA controls, ranked by mean differential accessibility. Transcription factor pairs with significant changes in accessibility owing to transcription factor order are colored. Transcription factor orders with significant differential accessibility between DE cells and ESCs are starred (significance computed by paired t-test and Wilcoxon signed-rank with Benjamini–Hochberg correction at FDR < 0.05).











