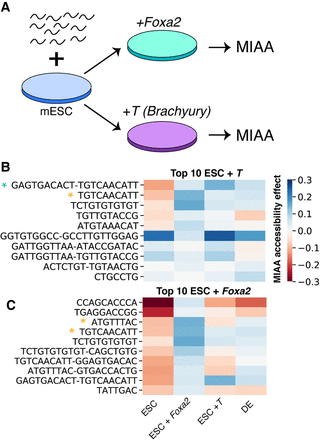
Overexpression of DE lineage-defining transcription factors results in changes to certain motifs representing DNA binding. (A) Synthetic DNA sequence library is integrated into ESCs, and Foxa2 and T are overexpressed. (B) Regression weight heatmap of top motifs and motif pairs that increase accessibility under T overexpression compared with ESCs. Blue star indicates motif visually matches T homodimer in ± orientation that is enriched in ChIP-seq peaks. Yellow star indicates motif is statistically enriched in ChIP-seq peaks of T binding in mouse definitive endoderm cells (P < 0.05 HOMER motif enrichment with Benjamini–Hochberg correction). (C) Regression weight heatmap of top motifs and motif pairs that increase accessibility under Foxa2 overexpression compared to ESCs. Star indicates motif is statistically enriched in FOXA2 ChIP-seq peaks in mouse DE cells (P < 0.05 HOMER motif enrichment with Benjamini–Hochberg correction).











