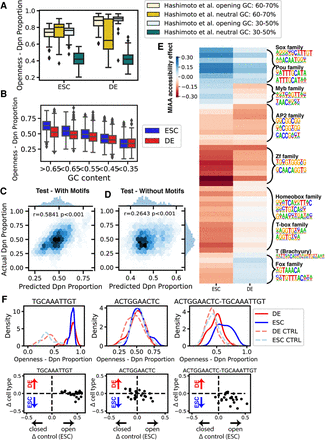
MIAA identifies global influence of GC-content and differentially accessible motifs. (A) GC-content observed to be correlated with accessibility in both stem and endoderm cells from positive (Hashimoto et al. opening) and negative (Hashimoto et al. neutral) control sequences. (B) GC-content correlated with accessibility in random DNA sequences. A regression model was trained on MIAA Dpn proportions with GC-content, replicate, and cell type–specific effects of 20 motifs and 26 motif pairs as features, and predicts well on (C) held-out test data (n = 4404) and performs significantly better than (D) a model trained without motif variables (adjusted R-squared motif model = 0.398; adjusted R-squared no motif model = 0.095). The correlation reported is the Pearson correlation coefficient (r). (E) Regression weights of individual motifs and motif pairs in stem and DE cells. Hierarchical clustering of regression weights followed by motif enrichment recovers clusters representing cell type–specific transcription factor DNA-binding motifs. (F) Example of individual motifs (left, middle) that alone do not result in differentially open chromatin but result in differentially open chromatin ESCs in combination (right). (Top row) Distribution of MIAA-measured accessibility in ESCs and DE cells for KMAC- or DeepAccess-generated motif, tested over 24 neutral sequence backgrounds and randomly shuffled DNA controls (CTRL). (Bottom row) Measurements for a particular DeepAccess or KMAC motif, in which each dot represents a single neutral background. The y-axis is the difference between endoderm and ESC accessibility, and the x-axis is the difference between each DNA sequence and its shuffled control in ESCs.











