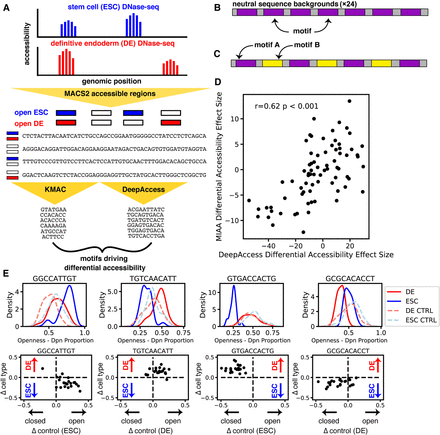
Differentially accessible motif generation from DNase-seq data validated by MIAA. (A) DNase-seq accessible regions called with MACS2 and 100-nt sequences extracted centered at narrow peak. KMAC and DeepAccess were applied to extract significant motifs potentially driving differential accessibility between ESCs and endoderm. (B) DNA sequences were designed using seven instances of each motif at the same locations in each DNA sequence inserted into 24 100-nt neutral sequence backgrounds, as well as pairs of motifs (C). (D) Predictions from DeepAccess for differential accessibility replicate experimental results (effect size by paired t-test between ESC and DE measurements). The correlation reported is the Pearson correlation coefficient (r). (E) Motif sequences show differential accessibility via opening ESC, opening endoderm, closing ESC, and closing endoderm (left to right). (Top row) Distribution of MIAA-measured accessibility in ESCs and DE cells for KMAC- or DeepAccess-generated motif, tested over 24 neutral sequence backgrounds and randomly shuffled DNA controls (CTRL). (Bottom row) Measurements for a particular DeepAccess or KMAC motif. Each dot represents a single neutral background. The y-axis is the difference between endoderm and ESC accessibility, and the x-axis is the difference between each DNA sequence and its shuffled control. The cell type in which control measurement is made is in parentheses.











