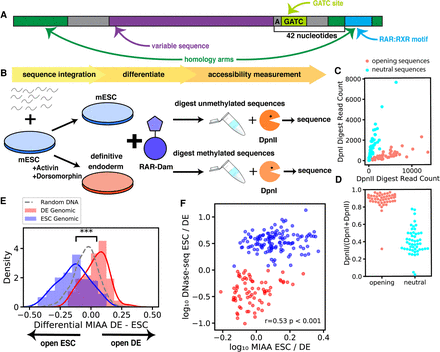
Multiplexed integrated accessibility assay (MIAA) measures local DNA accessibility of synthesized oligonucleotide DNA sequence libraries. (A) The MIAA library sequence construct contains a variable DNA sequence, homology arms for CRISPR-mediated HDR integration at a specific genomic locus that includes a binding site for retinoic acid receptor 42 nt downstream from the variable DNA sequence, and GATC site for DNA adenine methylase (Dam) methylation 1 nt downstream from the variable DNA sequence. (B) DNA sequences of 150 nt are integrated into ESCs at a designated genomic locus. ESCs are split, and half are differentiated into DE cells. Retinoic acid receptor fused to hyperactivated Dam enzyme results in methylation of DNA sequences that open DNA. DNA is extracted, and half is exposed to DpnII, which cleaves unmethylated sequences, whereas half is exposed to DpnI, which cleaves methylated sequences. Sequences are PCR amplified and sequenced. (C) DpnI and DpnII read counts measured from a single DE replicate show difference between designed chromatin opening and neutral DNA sequences. (D) Proportion of DpnII read counts measured from a single ESC replicate gives estimate of MIAA openness. (E) Genomic sequences are differentially DE accessible or ESC accessible as reported by difference between MIAA Dpn proportion in definitive endoderm compared with ESCs with randomly shuffled DNA control sequences (significance computed by Wilcoxon rank-sum test). (F) Differential accessibility as measured by log change in normalized DNase-seq reads and MIAA methylation proportion shows correlation between native differential accessibility and MIAA accessibility. The correlation reported is the Pearson's correlation coefficient (r).











