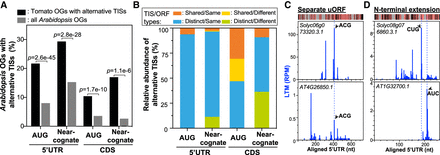
The conservation of alternative TISs between tomato and Arabidopsis. (A) The proportion (%) of Arabidopsis orthologous genes (OGs) with alternative TISs when the tomato OG has an alternative TIS (black; n = 1320, 1089, 473, and 65, from left to right) and the proportion (%) of all Arabidopsis OGs with alternative TISs (gray; n = 9349), which were considered as a background data set. The alternative TISs were categorized into four groups depending on their location (5′ UTR or CDS) and the type of initiation codon (AUG or near-cognate codon). P-values are the test of whether the proportion of Arabidopsis OGs with alternative TISs differs between the black and gray data sets (Fisher's exact test). (B) The relative proportion (%) of alternative TISs detected in both genes of a given tomato-Arabidopsis OG pair. Pairs were categorized into four groups depending on (1) whether the TIS position and codon are identical between orthologs (i.e., “Shared”) or different (i.e., “Distinct”) and (2) whether the corresponding ORF type between orthologs is the same (i.e., “Same”) or different (i.e., “Different”) (Supplemental Fig. S9; Methods). (C) Plots of LTM read density along the aligned 5′ UTRs of an OG pair with alternative TISs (arrow) at the same position and corresponding to the same ORF type. The TIS codon and corresponding ORF type for each OG pair are shown beside the TIS peak and on the top left, respectively. The heatmap (top) shows the conserved (pink) and nonconserved (black) sequences and gapped regions (gray). (D) As defined in C, but for the OG pairs with alternative TISs located at different positions but producing the same ORF type.











