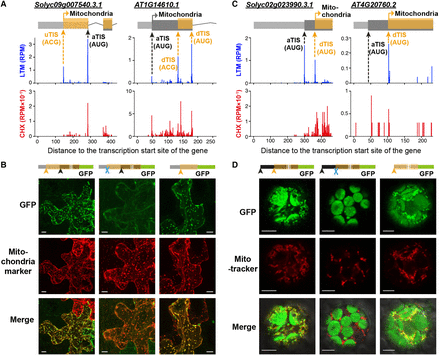
The differential localization of proteins encoded by genes with alternative in-frame TISs. (A,C) As indicated in Figure 1D, but for the orthologous gene pairs Solyc09g007540 and AT1G14610 (A) and Solyc02g023990 and AT4G20760 (C), which have identified alternative TISs (orange arrows) in frame with the annotated TIS (aTIS) that are either located upstream (uTIS) or downstream (dTIS) and encode N-terminally extended or truncated protein isoforms. The TIS of the protein isoform with predicted mitochondria targeting signals is indicated. See Supplemental Fig. S3 for full-length gene models. (B) Confocal images showing the localization of Solyc09g007540-GFP proteins with translation driven by the wild-type 5′ UTR and CDS (left), uTIS-mutated 5′ UTR and CDS (middle), and 5′ UTR region (right) in transiently transformed tobacco leaves. (Scale bar) 10 µm. The aTIS, uTISs/dTISs, and mutated TISs are indicated by a black arrow, orange arrow, and blue cross, respectively. (Mitochondria marker) CD3-992 (Nelson et al. 2007). (D) As described in B, but with the localization of Solyc02g023990-GFP proteins driven by the wild-type full-length CDS (left), dTIS-mutated full-length CDS (middle), and CDS region downstream of the dTIS (right) in transiently transformed Arabidopsis protoplasts. (MitoTracker) a mitochondria dye. (Scale bar) 5 µm.











