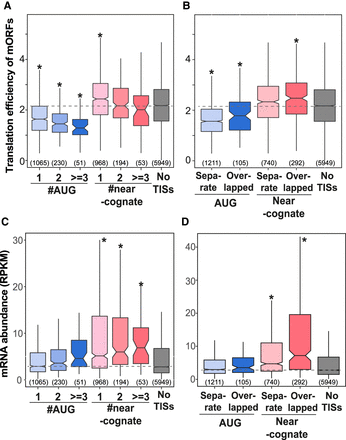
The correlation between alternative AUG/near-cognate TISs and translation efficiency/transcript abundance. (A) The translation efficiencies of main ORFs (mORFs) for genes with the indicated numbers of exclusively AUG-initiated TISs, exclusively near-cognate-initiated TISs, and without AUG/near-cognate-initiated TISs (“No TIS”) in the 5′ UTR. (B) The translation efficiencies of mORFs for genes with upstream AUG-initiated or near-cognate-initiated ORFs that are fully separate from or overlap and are out of frame with mORFs. (C,D) As described in A and B, but for steady-state mRNA abundances of genes. (*) P < 1 × 10−4. P-values are the test of whether the translation efficiency of mORFs and mRNA abundance determined for genes with AUG or near-cognate TISs differ from those of “No TIS” genes (Mann–Whitney U test). Dashed line shows the median value from the data for “No TIS” genes. Only genes with AUG or near-cognate sites in the 5′ UTR and with RPKM values ≥1 in the CDS region in both the CHX and mRNA RNA-seq data sets were included. The number of genes within a given group is shown in parentheses.











