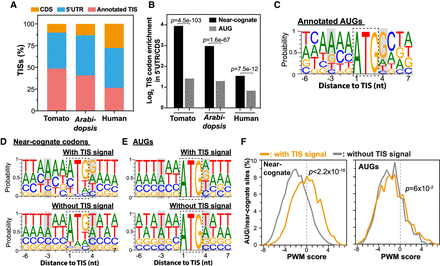
Characteristics of alternative AUG and near-cognate start codons. (A) The relative distribution (%) of the identified TISs located in annotated TISs (pink), 5′ UTRs (blue), and CDSs (orange) in tomato, Arabidopsis, and humans. (B) The differential distribution of AUG (gray) and near-cognate (black) TISs between 5′ UTRs and CDSs in tomato, Arabidopsis, and humans. P-values are the statistical significance test of whether the enrichment of near-cognate codons between 5′ UTRs and CDSs differs from that of AUGs (Fisher's exact test). (C) The probability of occurrence of ATCG nucleotides in sequence regions around the annotated TISs in tomato. Gray boxes highlight the −3 and +4 positions of the Kozak sequence. (D,E) As in C, but for the upstream near-cognate (D) and AUG (E) codons with TIS signals (i.e., located at TISs, top) and without TIS signals (bottom). (F) Position-weight matrix (PWM) scores of codon sites with TIS signals (orange) and without TIS signals (gray) for near-cognate codons (left) and AUG codons (right). PWM score was used to represent the sequence similarity between the regions surrounding a given codon site and those surrounding annotated TISs (Methods). P-values are the test of whether the PWM scores generated based on the codon sites with TIS signals (orange) differ from those of the codon sites without TIS signals (gray) (Mann–Whitney U test).











