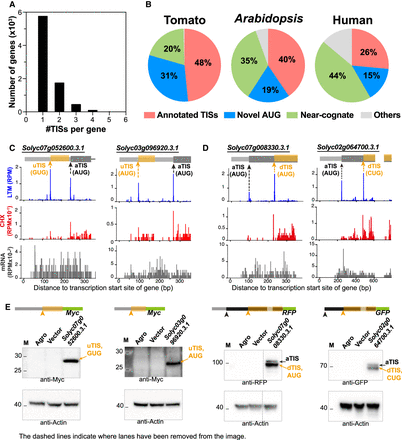
Alternative initiation of translation at AUG and near-cognate codons in vivo. (A) Distribution of the number of identified in vivo TISs per gene in tomato. (B) Pie charts showing the proportion (%) of in vivo TISs in tomato (n = 11,488), Arabidopsis (n = 11,909), and humans (n = 7974) that are annotated TISs (pink) or novel TISs containing an AUG (blue), near-cognate (green), or other codons (gray). (C,D) Plots of read densities across genes as described in Figure 1D, but for genes with alternative in-frame (left) and out-of-frame (right) TISs in the 5′ UTR that initiate separate ORFs (C) and with alternative in-frame TISs in the CDS that lead to N-terminally truncated ORFs (D). Within the gene models (top), the orange arrows indicate upstream and downstream TISs (uTIS/dTIS) located in the 5′ UTR (light gray boxes) and within annotated CDSs (dark gray boxes), respectively, and the orange boxes are the alternative TIS-initiated ORFs. Examples of translation initiating at AUG, GUG, or CUG are shown. See Supplemental Fig. S3 for full-length gene models. (E) Immunoblotting analysis of proteins encoded by transcripts with annotated and/or alternative TISs indicated in C and D and transiently expressed in tobacco leaves. (Agro) tobacco leaves infiltrated with agrobacteria without expression plasmid; (Vector) tobacco leaves infiltrated with agrobacteria containing the expression vector (i.e., the plasmid without target gene sequences).











