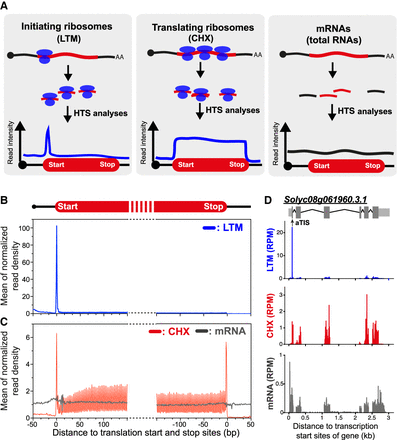
Comparison of the densities of LTM- and CHX-associated ribosome-protected fragments (RPFs) in genes. (A) An illustration of high-throughput sequencing (HTS) read distribution on mRNAs from CHX- and LTM-treated samples, which induced ribosome stalling, and untreated (total mRNA) samples. (B,C) Metagene plots of mean normalized read densities in regions around translation start and stop sites of genes calculated for LTM-treated (B) and CHX-treated and untreated (mRNA) samples (C). Normalized read density of a gene was calculated by normalizing the read count per base to the average read density for the entire CDS. Only genes with 5′ UTRs and 3′ UTRs ≥ 20 nt were included. (D) As described in B and C, but for a single gene, Solyc08g061960, with a TIS peak identified at an annotated TIS (aTIS). (RPM) reads per million mapped reads. Within the gene model (top), light gray boxes indicate UTRs, dark gray boxes indicate annotated CDSs, thin lines indicate introns, and black arrow indicates aTIS.











