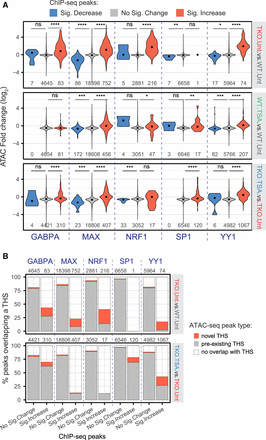
Transcription factor occupancy can promote chromatin accessibility in mESCs with perturbed DNA methylation or HDAC activity. (A) Quantitation of ATAC-seq fold change at transcription factor ChIP-seq peaks grouped according to their differential occupancy. Black dots indicate the median ATAC-seq fold change. One-tailed Mann–Whitney U tests; (*) P-value <0.05, (**) P-value <0.01, (***) P-value < 0.001, (****) P-value < 10−10, (ns) nonsignificant (P-value > 0.05). (B) Percentage of ChIP-seq peaks that overlap with a Tn5 hypersensitive site (THS). In the top panel, a “pre-existing” THS refers to an ATAC-seq peak identified in samples generated from untreated WT cells, whereas a novel THS is identified in DNMT.TKO but not WT cells. In the bottom panel, a “pre-existing” THS was identified in samples generated from untreated DNMT.TKO cells, whereas a novel THS is identified in TSA-treated DNMT.TKO cells only.











