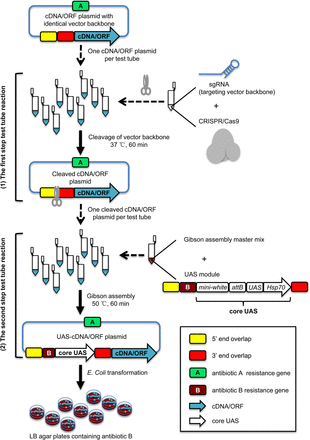
The CRISPRmass pipeline for construction of a UAS-cDNA/ORF library. (1) The first step test tube reaction. The identical vector backbones of cDNA/ORF plasmids are cleaved by Cas9/sgRNA. The cleavage site is located in vector backbones adjacent upstream of the cDNA/ORF 5′ end. The cleavage products, without purification, are directly subjected to the second step test tube reaction. The sgRNAs used in cleavage reactions are prepared by in vitro transcription and can be used directly without purification. Then, 28–40 bp of the 5′ end of the cleavage site (yellow box) is defined as “5′ end overlap,” and 28–40 bp of the 3′ end of the cleavage site (red box) is defined as “3′ end overlap.” The vector backbone carries the antibiotic A resistance gene (green box). (2) The second step test tube reaction. A vector-specific UAS module is joined into the Cas9-linearized cDNA/ORF plasmids right upstream of cDNA/ORF 5′ end through Gibson assembly, resulting in UAS-cDNA/ORF constructs. Gibson assembly products are directly subjected to E. coli transformation. Transformants are selected on LB agar plates containing antibiotic B corresponding to the antibiotic B resistance gene (brown box) of the vector-specific UAS module. Only the desired UAS-cDNA/ORF colonies can grow. A UAS module comprises 10 copies of UAS, a Hsp70 minimal promoter, an attB sequence for phiC31-mediated genomic integration, a mini-white transformation marker for Drosophila transgenesis, a selectable antibiotic B resistance gene for positive selection, and the 5′ end overlap and 3′ end overlap enabling Gibson assembly. Gibson assembly filters out any potential off-target DNA cleavages caused by CRISPR/Cas9.











