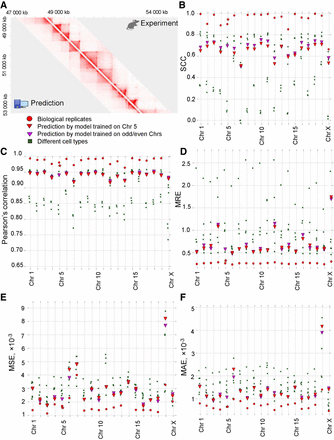
3DPredictor efficiently reconstructs spatial interactions based on CTCF occupancy, expression, and genomic distance. (A) Representative region of mouse Chromosome 2 showing predicted and experimentally derived Hi-C interactions in mouse hepatocytes. (B–F) Various metrics of 3DPredictor accuracy for each chromosome of mouse hepatocytes. Circles represent comparison between two replicas; green squares show comparisons between hepatocytes and other cell types. Red triangles display 3DPredictor results obtained using single Chromosome 5 for training; data obtained when validating on the same chromosome are marked with stars. Purple triangles show results of 3DPredictor trained on 10 chromosomes (results for even chromosomes obtained using model trained on odd chromosomes and vice versa).











