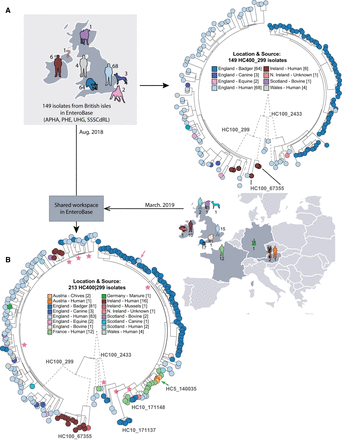
Effects of sample bias on inferred transmission chains within HC400_299 Agama isolates. (A, left) Map of hosts in the British Isles of 149 Agama isolates in EnteroBase in August, 2018. (Right) Maximum-likelihood radial phylogeny (http://enterobase.warwick.ac.uk/a/21773/d) based on RAxML (Stamatakis 2014) of 8791 nonrepetitive core SNPs as calculated by EnteroBase Dendrogram against reference genome 283179. Color coding is according to a user-defined field (Location & Source). HC100 cluster designations for three microclades are indicated. HC100_2433 contained all Agama from badgers. (B, right) Summary of hosts and countries from which 64 additional Agama isolates had been sequenced by March 2019. (Left) Maximum-likelihood radial dendrogram (http://enterobase.warwick.ac.uk/a/23882/d) based on 9701 SNPs from 213 isolates. Multiple isolates of Agama in HC100_2433 were now from humans and food in France and Austria. HC100_299 and HC100_67355 now contained multiple isolates from badgers, livestock, companion animals, and mussels, demonstrating that the prior strong association of Agama with humans and badgers in A reflected sample bias. Stars indicate multiple MRCAs of Agama in English badgers, whereas the pink arrow indicates a potential transmission from badgers to a human in Bath/North East Somerset, which is close to Woodchester Park. The green arrow indicates a potential food-borne transmission chain consisting of four closely related Agama isolates in HC5_140035 from Austria (chives × 2; human blood culture × 1) and France (human × 1) that were isolated in 2018. The geographical locations of the badger isolates are shown in Supplemental Figure S5.











