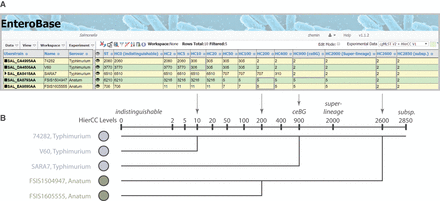
The hierarchical cgMLST clustering (HierCC) scheme in EnteroBase. (A) A screenshot of Salmonella cgMLST V2 plus HierCC V1 data for five randomly selected genomes. The numbers in the columns are the HierCC cluster numbers. Cluster numbers are the smallest cgMLST ST number in single-linkage clusters of pairs of STs that are joined by up to the specified maximum number of allelic differences. These maximum differences are indicated by the suffix of each HC column, starting with HC0 for 0 cgMLST allelic differences, other than missing data, through to HC2850 for 2850 allelic differences. The cluster assignments are greedy because individual nodes which are equidistant from multiple clusters are assigned to the cluster with the smallest cluster number. (B) Interpretation of HierCC numbers. The assignments of genomic cgMLST STs to HC levels can be used to assess their genomic relatedness. The top two genomes are both assigned to HC10_306, which indicates a very close relationship, and may represent a transmission chain. The top three genomes are all assigned to HC900_2, which corresponds to a legacy MLST eBG. HC2000 marks superlineages (Zhou et al. 2018c), and HC2850 marks subspecies. This figure illustrates these interpretations in the form of a cladogram drawn by hand.











