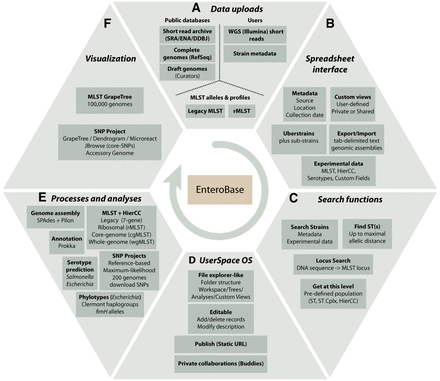
Overview of EnteroBase Features. (A) Data uploads. Data are imported from public databases, user uploads, and existing legacy MLST and rMLST databases at PubMLST (https://pubmlst.org/). (B) Spreadsheet Interface. The browser-based interface visualizes sets of strains (one Uberstrain plus any number of substrains) each containing metadata, and their associated experimental data and custom views. Post-release data can be exported (downloaded) as genome assemblies or tab-delimited text files containing metadata and experimental data. Metadata can be imported to entries for which the user has editing rights by uploading tab-delimited text files. (C) Search Strains supports flexible (AND/OR) combinations of metadata and experimental data for identifying entries to load into the spreadsheet. Find ST(s) retrieves STs that differ from a given ST by no more than a maximal number of differing alleles. Locus Search uses BLASTN (Altschul et al. 1990) and UBlastP in USEARCH (Edgar 2010) to identify the MLST locus designations corresponding to an input sequence. Get at this level: menu item after right clicking on experimental MLST ST or cluster numbers. (D) UserSpace OS. A file explorer–like interface for manipulations of workspaces, trees, SNP projects, and custom views. These objects are initially private to their creator but can be shared with buddies or rendered globally accessible. (E) Processes and analyses. EnteroBase uses EToKi and external programs as described in Supplemental Figure S1. (F) Visualization. MLST trees are visualized with the EnteroBase tools GrapeTree (Zhou et al. 2018a) and Dendrogram, which in turn can transfer data to external websites such as Microreact (Argimón et al. 2016).











