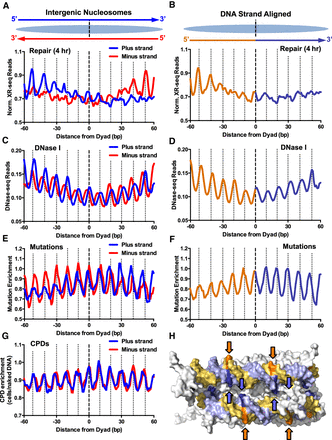
Asymmetric repair of human intergenic nucleosomes promotes a DNA strand polarity in somatic mutations in melanoma. (A) Normalized NER activity is elevated on the 5′ side of both DNA strands in intergenic nucleosomes. Normalized XR-seq reads for the 4-h repair time point were plotted for both DNA strands in their normal antiparallel orientation (top) at positions from −60 to +60 bp from the dyad axis of intragenic nucleosomes. (B) Same as A, except the plus and minus DNA strands were both aligned in the 5′-to-3′ direction, and the average of both aligned DNA strands is plotted. Data for the 5′ side of the dyad axis are depicted in orange, and data for the 3′ side of the dyad are depicted in purple. (C,D) Same as A and B, except DNase-seq read density (Degner et al. 2012) was plotted. (E,F) Same as A and B, except mutation enrichment in melanoma tumors was plotted. (G) Same as A, except CPD enrichment was plotted. (H) Structural model showing that at “out” positions (indicated with arrows), the 5′ side of the nucleosomal DNA (orange/gold) faces the solvent, whereas the 3′ side of the nucleosomal DNA (light purple/blue) faces the other DNA gyre, and is thereby less accessible for repair. PyMol (https://pymol.org/2/) was used to visualize the nucleosome structure (PDB ID: 1KX5) from Davey et al. (2002).











