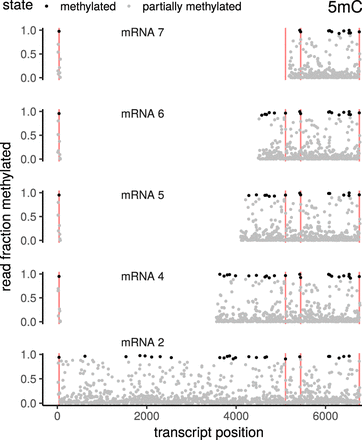
5mC methylation of various annotated coronavirus mRNAs. The transcripts have been aligned such that corresponding genomic positions can be found in the same vertical column across facets. Both the leader sequence (including the TRS) and the nested sg sequences show consistent patterns of methylation across transcripts. Exemplary positions that display consistent methylation across all investigated transcripts are marked as red vertical lines. Note that although coronavirus recombination uses two TRSs, the resulting transcript has only one TRS, because of self-similarity-based pairing of the TRS. Positions have been labeled “methylated” if at least 90% of the reads show a methylation signal. Using this threshold, the estimated FPR is <5%.











