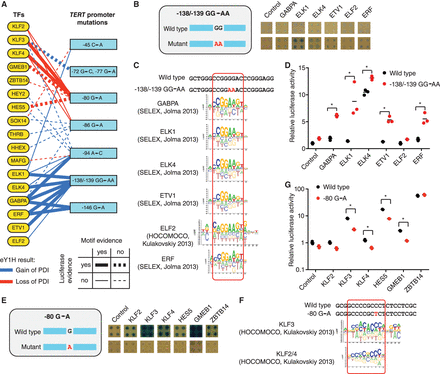
Identification of altered TF binding to single- and di-nucleotide mutations in the TERT promoter. (A) Gain and loss of protein-DNA interactions (PDIs) with seven TERT single- and di-nucleotide mutations in the TERT promoter were identified by eY1H assays. Mutation position is relative to the initiation codon. Blue line: gain of PDIs, red line: loss of PDIs. Full lines indicate validation by motif analysis. Thick lines indicate validation by luciferase assays. (B,E) eY1H screen for wild type and a −138/−139 GG→AA (B) or a −80 G→A (E) mutation in the TERT promoter associated with cancer. Each interaction was tested in quadruplicate. Control: empty AD-vector. (C,F) Motifs obtained from Cis-BP that match the differential TFs identified by eY1H assays. (D,G) Luciferase assays to validate the differential TF interactions with the TERT promoter alleles. Relative luciferase activity is plotted as fold change compared to cells cotransfected with the wild-type TERT promoter construct and the VP160 vector (control). A representative experiment of three is shown. The average of three replicates is indicated by the black line. (*) P < 0.05 by one-tailed log-transformed Student's t-test with Benjamini-Hochberg correction.











