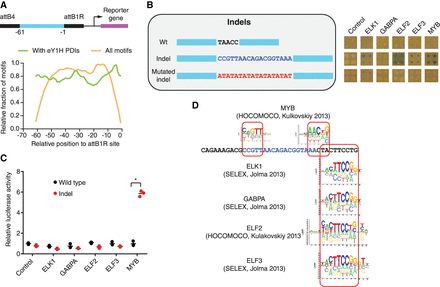
Identification of altered TF binding to a TAL1 superenhancer insertion. (A) Relative fraction of motifs spanning the indicated position relative to the attB1R site (orange line) and the relative fraction of motifs for which eY1H interactions were detected (green line) for 168 sequences of 61 bp tested by eY1H assays. Sliding windows of 3 bp were used. (B) eY1H screen for an 18-bp insertion in a TAL1 superenhancer associated with T cell acute lymphoblastic leukemia. Wild type and an (AT)9 sequence replacement (for the 18-bp insertion) were screened as controls. Each interaction was tested in quadruplicate. Control: empty AD-vector. (C) Luciferase assays to validate the differential TF interactions with the TAL1 superenhancer wild-type and insertion alleles. HEK293T cells were cotransfected with reporter plasmids containing the wild-type or insertion TAL1 superenhancer region cloned upstream of the firefly luciferase reporter gene and expression vectors for the indicated TFs (fused to the activation domain VP160). After 48 h, cells were harvested and luciferase assays were performed. Relative luciferase activity is plotted as fold change compared to cells cotransfected with the wild-type TAL1 superenhancer construct and the VP160 vector (control). A representative experiment of three is shown. The average of three replicates is indicated by the black line. (*) P < 0.05 by one-tailed log-transformed Student's t-test with Benjamini-Hochberg correction. (D) Motifs obtained from Cis-BP that match the differential TFs identified by eY1H assays.











