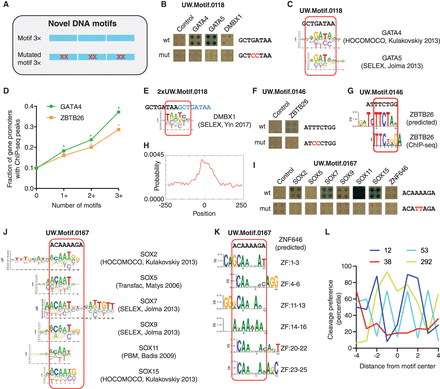
Identification of TFs that bind to novel DNA motifs. (A) Motifs were tested by eY1H assays as three tandem copies. Motifs with two mutated bases were tested as controls. (B,F,I) eY1H screen against 1086 TFs of three motifs identified by DNase I footprinting by the ENCODE Project. Tested sequences are indicated. Each interaction was tested in quadruplicate. Control: empty AD-vector. (C,E,J) Alignment of motif logos from Cis-BP to the tested sequences. (D) Fraction of genes with ChIP-seq peaks for GATA4 and ZBTB26 in their promoters as a function of the number of UW.Motif.0118 and UW.Motif.0146 motifs, respectively. (*) P < 0.001 versus absence of motif by proportion comparison test. (G) Predicted ZBTB26 motif based on the amino acid sequence of zinc fingers 1–3 and highest scoring motif derived from ChIP-seq data. (H) Location of the ZBTB26 motif derived from ChIP-seq data within the peaks. (K) Predicted ZNF646 motif based on the amino acid sequence of zinc fingers 1–3, 4–6, 11–13, 14–16, 20–22, or 23–25. (L) Percentile cleavage preference by DNase I at each position for UW.Motif.0012, UW.Motif.0038, UW.Motif.0053, and UW.Motif.0292.











