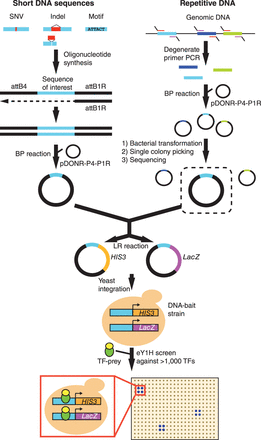
eY1H assays to test short DNA sequences and repetitive DNA elements. Short DNA sequences are synthesized flanked by the attB4 and attB1R Gateway sites. The reverse strand is synthesized by primer extension (attB1R). The double-stranded DNA generated is cloned into the pDONR-P4-P1R vector by Gateway BP reaction. Repetitive DNA sequences are amplified from genomic DNA using degenerate primers flanked by the attB4 (forward) or the attB1R (reverse) sites. The repetitive element DNA library generated is cloned en masse into the pDONR-P4-P1R vector, and individual sequences are selected after bacterial transformation and picking of individual colonies. Both short DNA sequences and repetitive DNA are then transferred into eY1H reporter vectors (HIS3 and LacZ) and integrated into the yeast genome to generate chromatinized DNA-bait strains. DNA-bait strains are tested for interactions against an array of 1086 human TF-preys (TFs fused to the yeast Gal4 activation domain) by mating. Interactions are identified by the ability of yeast colonies to grow in the absence of histidine and in the presence of the His3p inhibitor 3-amino-1,2,4-triazole and to turn blue in the presence of the β-galactosidase substrate X-gal. Interactions are tested in quadruplicate.











