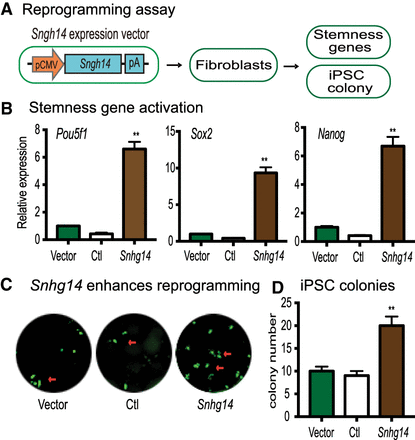
Snhg14 lncRNA enhances pluripotent reprogramming. (A) Schematic diagram of the reprogramming assay. Fibroblasts and MEF cells were transfected with Snhg14 lentivirus. After puromycin selection, the Snhg14-expressing cells were collected for quantitative PCR measurement of the endogenous stemness genes (Pou5f1, Sox2, and Nanog) or for reprogramming. (B) Activation of the endogenous stemness genes by Snhg14. Expression of Pou5f1, Sox2, and Nanog was measured by quantitative PCR and calculated as relative expression by setting the Vector control as 1. (**) P < 0.01 as compared with the Ctl and Vector controls. (C) Snhg14 enhances the efficiency of reprogramming. MEF cells were transfected with the lentiviruses carrying the empty vector (Vector), lncRNA control (Ctl), and Snhg14. After doxycycline (DOX) induction, the iPSC colonies were immunostained using anti-NANOG antibody (green). (D) Quantitation of iPSC colonies. iPSC colonies per well were counted and averaged from three independent assays. (**) P < 0.01 as compared with the Ctl and Vector controls.











