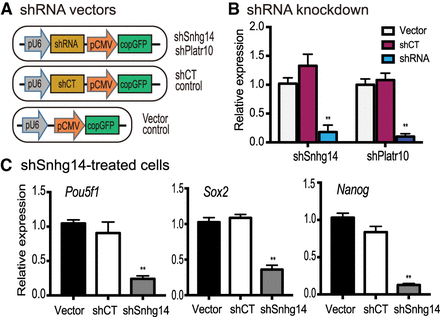
Knockdown of Sox2-interacting lncRNAs down-regulates stemness genes. (A) Lentiviral shRNA knockdown vectors. (pU6) RNA polymerase III U6 promoter; (pCMV) CMV promoter; (CopGFP) fluorescence tracking marker; (shRNA) shRNAs targeting Snhg14 and Platr10; (shCT) shRNA random control. Lentiviruses were packaged in 293T cells and were used to infect iPSCs. (B) Knockdown of Snhg14 and Platr10 lncRNAs in iPSCs. After lentiviral transfection, the shRNA-expressing iPSCs were selected by puromycin and were collected for quantitative PCR. (**) P < 0.01 as compared with shCT and vector controls. (C) Down-regulation of three core stem cell factor genes in Snhg14-knocking down iPSCs. (**) P < 0.01 as compared with shCT and vector controls.











