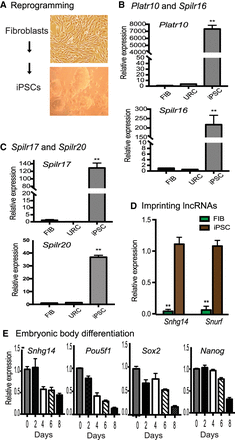
Sox2-interacting lncRNAs are associated with reprogramming. (A) Schematic diagram of pluripotent reprogramming. Fibroblasts and iPSCs collected at different stages of reprogramming. (B) Differential expression of the Sox2-binding lncRNAs (Platr10 and Spilr16) in reprogramming. (Fib) Fibroblasts; (URC) unreprogrammed cells that express the OSKM factors, but fail to complete reprogramming; (iPSC) induced pluripotent stem cells; (Spilr) Sox2 promoter-interacting long noncoding RNA. The data shown are mean ± SEM from three independent experiments. (**) P < 0.01 as compared with fibroblasts and unreprogrammed cells. (C) Differential expression of Spilr17 and Spilr20. (**) P < 0.01 as compared with fibroblasts and unreprogrammed cells. (D) Quantitative PCR of imprinted Snurf and Snhg14 in reprogramming. (**) P < 0.01 as compared with iPSCs. (E) Dynamic expression of Snhg14 in embryoid body differentiation. iPSCs were collected at different stages of embryoid body formation and used for quantitative PCR. Note the similar expression pattern of Snhg14 to the stem cell marker genes.











