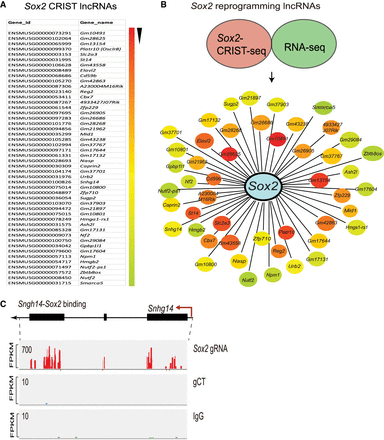
The lncRNA interacting network in the Sox2 promoter. (A) CRIST-seq identifies the top 50 Sox2 promoter-interacting RNAs. The Sox2 interacting RNAs are listed in order of the enrichment fold of the top 50 CRIST-seq data. (B) The reprogramming-associated Sox2 lncRNA interacting network. To identify reprogramming-associated lncRNAs, fibroblasts and iPSCs were collected at different stages of reprogramming, and total RNAs were sequenced. Data regarding the RNAs that were changed by greater than twofold were combined with the CRIST-seq data using a VENN program. A cut-off threshold of peak enrichment FPKM > 50 was arbitrarily set to select CRIST-seq RNAs for VENN analysis. Integration of these two data sets generated a total of 59 RNAs, which were differentially expressed in reprogramming and also interacted with the Sox2 promoter. The Sox2 RNA interaction was drawn based on the differential expression fold (red to blue) of lncRNAs between iPSCs and fibroblasts. (C) Specific binding of reprogramming-associated lncRNA Snhg14 in the Sox2 promoter chromatin complex. Three sets of CRIST-seq BAM data (Sox2-gRNA, control gCT, and IgG control) were uploaded onto the Integrative Genomics Viewer (IGV) browser (Thorvaldsdóttir et al. 2013), and a Sashimi plot was used to compare the enrichment signal between each group. (FPKM) Fragments per kilobase of exon per million fragments mapped. The CRIST-seq data revealed that all three exons of Snhg14 interact with the Sox2 promoter.











