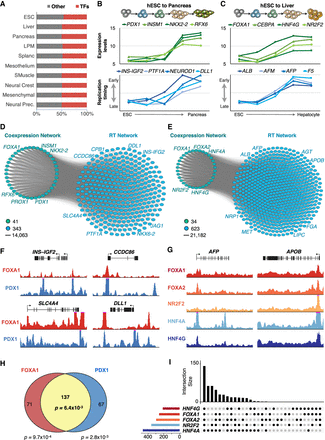
Bipartite networks. (A) Transcription factor expression patterns distinguish each cell type/differentiation intermediate. Cell-type–specific expression patterns were analyzed to identify the most significant differentially coexpressed genes, and then the resulting genes were classified as TFs or other type. (B) Expression patterns of exemplary key TFs of pancreas development correlate with the RT of downstream regulators of pancreatic differentiation. (C) Expression patterns of exemplary key TFs of liver development correlate with the RT of downstream regulators of hepatic differentiation. (D,E) Bipartite networks of pancreas (D) and liver (E) differentiation. Bipartite networks were constructed based on the correlation between transcriptional levels of the genes in the TRN side and RT changes of genes in the RT side. TRNs were constructed from the top genes coexpressed in pancreas and liver, respectively. RT network sides were constructed by identification of genes with RT patterns highly correlated with the expression levels of genes at the TRN side. Only edges between TRN and RT networks are shown (with each gene in the RT side connected with at least 50% of the nodes in the TRN side), as all nodes within each network are interconnected with all others. (F) ChIP-seq signals show the co-occupancy of FOXA1 and PDX1 at exemplary pancreatic-specific downstream targets predicted by the bipartite network shown in D. (G) ChIP-seq signals show the co-occupancy of FOXA1, FOXA2, NR2F2, HNF4A, and HNF4G at hepatic-specific downstream targets predicted by the bipartite networks shown in E. (H,I) Co-occupancy analysis for the pancreatic-specific and hepatic-specific TFs at the promoters of downstream targets.











