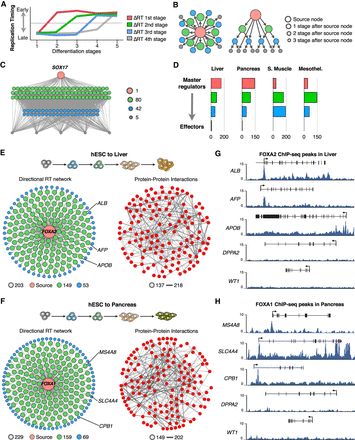
Directional RT networks. (A) Distinct genes change RT at different time points during cell fate commitment, and the order of RT changes can be used to construct directional RT networks. Red genes change during the first transition between differentiation stages, whereas gray genes change at the last differentiation transition. (B) 2D maps and hierarchical displays of a directional RT networks were constructed based on a source gene (central node) and downstream connected nodes. (C) An exemplary directional RT network for liver differentiation is shown. The central node is SOX17, and all downstream nodes were connected based on temporal times during differentiation at which they change RT. (D) Classification of gene nodes according to their hierarchy in directed RT networks. Master regulators were operationally defined as those genes that change RT in the earliest differentiation transition and have the largest degree of connectivity (red nodes). Green and blue nodes represent “manager” nodes that are connected to the final “effector” nodes at the lowest level of the network (gray nodes). Node distribution in each of the hierarchical levels for each differentiation pathway is shown. (E,F) Directed RT networks of FOXA2 and FOXA1 during liver and pancreatic differentiation, respectively. FOXA2 and FOXA1 (red node) change from late to early replication during the earliest stages of differentiation; potential downstream genes were identified as those genes that change in subsequent differentiation stages (green and blue nodes). Sixty-five percent or more of the genes within the directed RT networks interact with each other at the protein level (protein–protein interactions were obtained from the STRING database; nodes were mapped using the same layout as the directional RT network; and edge thickness indicates the strength of the data supporting each interaction). (G,H) ChIP-seq signals show the binding of FOXA2 and FOXA1 at the promoters of predicted hepatic-specific and pancreas-specific downstream genes (ALB, AFP, and APOB for liver and CPB1, MS4A8, and SLC4A4 for pancreas). A pluripotency gene (DPPA2) and a mesodermal-specific gene (WT1) are shown as a control.











