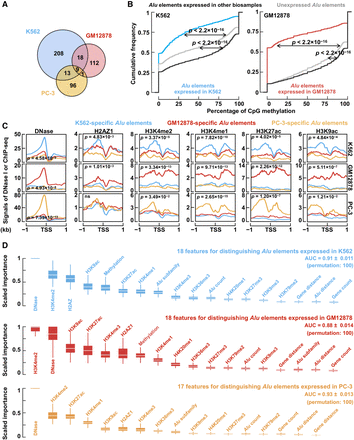
Cell-type–specific Alu expression corresponds to cell-type–specific histone modifications. (A) Cell-type–specific Alu expression in K562 (blue), GM12878 (red), and PC-3 (yellow) cells. (B) DNA methylation profile of expressed and unexpressed Alu elements. Wilcoxon rank-sum test P-values are shown. (C) Average signal of DNase-seq as well as ChIP-seq signals of H2AZ1, H3K4me2, H3K4me1, H3K27ac, and H3K9ac per cell type (rows) in the ±1-kb regions centered on the TSSs of Alu elements specifically expressed in each of the three cell types. Student's t-test P-values are shown for comparing the average signals in the corresponding cell type against the other two cell types (e.g., DNase signals in K562 cells for K562-specific Alu elements against DNase signals in K562 cells for GM12878-specific and PC-3-specific Alu elements). (D) Features in random forest models for distinguishing cell-type–specific Alu expression in K562 (blue), GM12878 (red), and PC-3 (yellow) cells against 150 randomly sampled Alu elements expressed in other cell types. The features are ordered by their importance.











